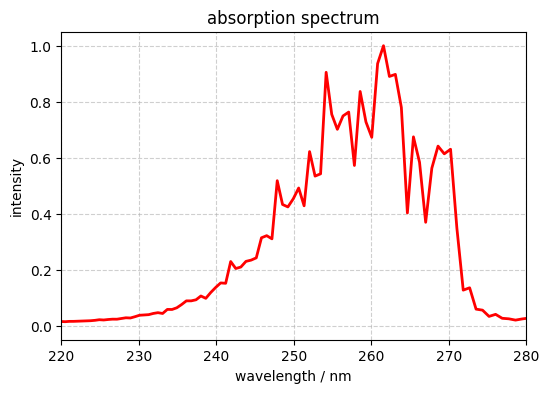
Example 16 Plot S2 absorption spectrum of pyrazine
References
Model Hamiltonian
The parameters in the Hamiltonian are given in electron volts (eV) while, in the calculation, these values were converted to atomic units (a.u.) . This model contains two excited states (S1 and S2).
\[\begin{split}\begin{align}
&\text{H}=\text{H}_{\text{el}}+\text{H}_{\text{vib}}+\text{H}_{G_{1}}+\text{H}_{G_{2}}+\text{H}_{G_{3}}+\text{H}_{G_{4}}
,
\\
&\text{H}_{\text{el}}=\begin{pmatrix} -0.4230 & 0 \\ 0 & 0.4230 \end{pmatrix}
,
\\
&\text{H}_{\text{vib}}=\sum_{i\in \{6a,1,9a...,16b,11\}}\frac{\omega_{i}}{2}(P_{i}^{2}+Q_{i}^{2})
,
\\
&\text{H}_{G_{1}}=
\sum_{i \in G_{1}}\underbrace{\begin{pmatrix} a_{i} & 0 \\ 0 & b_{i}\end{pmatrix}Q_{i}}_{\text{5 terms}}
,
\\
&\text{H}_{G_{2}}=
\sum_{(i,j) \in G_{2}}\underbrace{\begin{pmatrix}a_{ij} & 0 \\ 0 & b_{ij}\end{pmatrix}Q_{i}Q_{j}}_{\text{40 terms}}
,
\\
&\text{H}_{G_{3}}=
\sum_{i \in G_{3}}\underbrace{\begin{pmatrix} 0 & c_{i} \\ c_{i} & 0\end{pmatrix}Q_{i}}_{\text{1 term}}
,
\\
&\text{H}_{G_{4}}=
\sum_{(i,j) \in G_{4}}\underbrace{\begin{pmatrix}0 & c_{ij} \\ c_{ij} & 0\end{pmatrix}Q_{i}Q_{j}}_{\text{29 terms}}
\end{align}\end{split}\]
1.Import modules
[1]:
import platform
import sys
import pytdscf
print(sys.version)
print(f"pytdscf version = {pytdscf.__version__}")
print(platform.platform())
3.10.19 (main, Oct 9 2025, 15:25:03) [Clang 16.0.0 (clang-1600.0.26.6)]
pytdscf version = 1.3.0
macOS-26.2-arm64-arm-64bit
[2]:
import matplotlib.pyplot as plt
import netCDF4 as nc
import numpy as np
import sympy
from pympo import AssignManager, OpSite, SumOfProducts
from pytdscf import (
BasInfo,
Boson,
Exciton,
Model,
Simulator,
TensorHamiltonian,
TensorOperator,
units,
)
2.Set model parameters
[3]:
backend = "numpy"
# (eV)
# delta, omega coeff data : taken from https://pubs.aip.org/aip/jcp/article-abstract/110/2/936/475747
delta = 0.4230
omega = [
0.0739, 0.1258, 0.1525, 0.1961, 0.3788, # Ag (6a, 1, 9a, 8a, 2)
0.1139, # B1g (10a)
0.0937, 0.1219, # B2g (4, 5)
0.0873, 0.1669, 0.1891, 0.3769, # B3g (6b, 3, 8b, 7b)
0.0423, 0.1190, # Au (16a, 17a)
0.1266, 0.1408, 0.1840, 0.3734, # B1u (12, 18a, 19a, 13)
0.1318, 0.1425, 0.1756, 0.3798, # B2u (18b, 14, 19b, 20b)
0.0521, 0.0973 # B3u (16b, 11)
]
# data of G1 (eV)
# k_mode : [a_i, b_i]
g1_data = {
0: [-0.0981, 0.1355], # 6a
1: [-0.0503, -0.1710], # 1
2: [ 0.1452, 0.0375], # 9a
3: [-0.0445, 0.0168], # 8a
4: [ 0.0247, 0.0162] # 2
}
# data of G2 (eV)
# B1g x Ag
# except Ag x Ag
# (k_mode, l_mode) : [aij, bij]
g2_data = {
# Au modes (v16a=omega13, v17a=omega17)
(13, 13): [ 0.01145, -0.01459], # 16a * 16a
(17, 17): [-0.02040, -0.00618], # 17a * 17a
(13, 17): [ 0.00100, -0.00091], # 16a * 17a
# B1g mode (v10a:omega5)
(5, 5): [-0.01159, -0.01159], # 10a * 10a
# B2g modes (v4:omega6, v5:omega11)
(6, 6): [-0.02252, -0.03445], # 4 * 4
(11, 11): [-0.01825, -0.00265], # 5 * 5
(6, 11): [-0.00049, 0.00911], # 4 * 5
# B3g modes (v6b:omega7, v3:omega8, v8b:omega9, v7b:omega10)
(7, 7): [-0.00741, -0.00385], # 6b * 6b
(8, 8): [ 0.05183, 0.04842], # 3 * 3
(9, 9): [-0.05733, -0.06332], # 8b * 8b
(10, 10): [-0.00333, -0.00040], # 7b * 7b
(7, 8): [ 0.01321, -0.00661], # 6b * 3
(7, 9): [-0.00717, 0.00429], # 6b * 8b
(7, 10): [ 0.00515, -0.00246], # 6b * 7b
(8, 9): [-0.03942, -0.03034], # 3 * 8b
(8, 10): [ 0.00170, -0.00185], # 3 * 7b
(9, 10): [-0.00204, -0.00388], # 8b * 7b
# B1u modes (v12:omega12, v18a:omega14, v19a:omega15, v13:omega16)
(12, 12): [-0.04819, -0.00840], # 12 * 12
(14, 14): [-0.00792, 0.00429], # 18a * 18a
(15, 15): [-0.02429, -0.00734], # 19a * 19a
(16, 16): [-0.00492, 0.00346], # 13 * 13
(12, 14): [ 0.00525, 0.00536], # 12 * 18a
(12, 15): [-0.00485, -0.00097], # 12 * 19a
(12, 16): [-0.00326, 0.00034], # 12 * 13
(14, 15): [ 0.00852, 0.00209], # 18a * 19a
(14, 16): [ 0.00888, -0.00049], # 18a * 13
(15, 16): [-0.00443, 0.00346], # 19a * 13
# B2u modes (v18b:omega18, v14:omega20, v19b:omega21, v20b:omega22)
(18, 18): [-0.00277, -0.01179], # 18b * 18b
(20, 20): [ 0.03924, 0.04000], # 14 * 14
(21, 21): [ 0.00992, 0.01246], # 19b * 19b
(22, 22): [-0.00110, 0.00069], # 20b * 20b
(18, 20): [ 0.00016, -0.00844], # 18b * 14
(18, 21): [-0.00250, 0.07000], # 18b * 19b
(18, 22): [ 0.00357, -0.01249], # 18b * 20b
(20, 21): [-0.00197, -0.05000], # 14 * 19b
(20, 22): [-0.00355, 0.00265], # 14 * 20b
(21, 22): [ 0.00623, -0.00422], # 19b * 20b
# B3u modes (v16b:omega19, v11:omega23)
(19, 19): [-0.02176, -0.02214], # 16b * 16b
(23, 23): [ 0.00315, -0.00496], # 11 * 11
(19, 23): [-0.00624, -0.00261], # 16b * 11
}
# data of G3 (eV)
# mode 10a(omega5) : c_i
g3_val = 0.2080
# data of G4 (eV)
# (k_mode, l_mode) : c_ij
# v6a:omega0, v1:omega1, v9a:omega2, v8a:omega3, v2:omega4, v10a:omega5,
# v4:omega6, v6b:omega7, v3:omega8, v8b:omega9, v7b:omega10, v5:omega11,
# v12:omega12, v16a:omega13, v18a:omega14, v19a:omega15, v13:omega16, v17a:omega17,
# v18b:omega18, v16b:omega19, v14:omega20, v19b:omega21, v20b:omega22, v11:omega23
g4_data = {
# B1g x Ag
(5, 0): -0.01000, # 10a * 6a
(5, 1): -0.00551, # 10a * 1
(5, 2): 0.00127, # 10a * 9a
(5, 3): 0.00799, # 10a * 8a
(5, 4): -0.00512, # 10a * 2
# B2g x B3g
(6, 7): -0.01372, # 4 * 6b
(6, 8): -0.00466, # 4 * 3
(6, 9): 0.00329, # 4 * 8b
(6, 10):-0.00031, # 4 * 7b
(11, 7): 0.00598, # 5 * 6b
(11, 8):-0.00914, # 5 * 3
(11, 9): 0.00961, # 5 * 8b
(11, 10): 0.00500, # 5 * 7b
# Au x B1u
(13, 12): -0.01056, # 16a * 12
(13, 14): 0.00559, # 16a * 18a
(13, 15): 0.00401, # 16a * 19a
(13, 16): -0.00226, # 16a * 13
(17, 12): -0.01200, # 17a * 12
(17, 14): -0.00213, # 17a * 18a
(17, 15): 0.00328, # 17a * 19a
(17, 16): -0.00396, # 17a * 13
# B3u x B2u
(19, 18): 0.00118, # 16b * 18b
(19, 20): -0.00009, # 16b * 14
(19, 21): -0.00285, # 16b * 19b
(19, 22): -0.00095, # 16b * 20b
(23, 18): 0.01281, # 11 * 18b
(23, 20): -0.01780, # 11 * 14
(23, 21): 0.00134, # 11 * 19b
(23, 22): -0.00481 # 11 * 20b
}
[4]:
delta_sym = delta/ units.au_in_eV
omega_syms = [sympy.Symbol(f"omega_{i}") for i in range(len(omega))]
subs = {}
for omega_val, omega_sym in zip(
omega, omega_syms, strict=True
):
subs[omega_sym] = omega_val / units.au_in_eV
3.Setup basis for wavefunction
[5]:
nmode = len(omega)
nsite = (nmode + 1)
basis = []
for isite in range(nsite):
if isite == 0:
basis.append(Exciton(nstate=2))
else:
basis.append(Boson(nstate=10))
basinfo = BasInfo([basis])
4.Setup operator
[6]:
p = basis[1].get_p_matrix()
q = basis[1].get_q_matrix()
pp = basis[1].get_p2_matrix()
qq = basis[1].get_q2_matrix()
[7]:
q_ops = []
p_ops = []
qq_ops = []
pp_ops = []
for isite in range(nsite):
if isite == 0:
q_ops.append(None)
p_ops.append(None)
qq_ops.append(None)
pp_ops.append(None)
else:
q_ops.append(OpSite("Q_{" + f"{isite}" + "}", isite, value=q))
p_ops.append(OpSite("P_{" + f"{isite}" + "}", isite, value=p))
qq_ops.append(OpSite("Q2_{" + f"{isite}" + "}", isite, value=qq))
pp_ops.append(OpSite("P2_{" + f"{isite}" + "}", isite, value=pp))
[8]:
sop = SumOfProducts()
# H_el
el_mat = np.zeros((2, 2), dtype=np.float64)
el_mat[0,0]= -delta_sym
el_mat[1,1]= delta_sym
sop += OpSite("H_el", 0, value=el_mat)
# (eV) -> (au)
ev_to_au = 1.0 / units.au_in_eV
for k_mode in range(nmode):
jsite = 1 + k_mode
# H_vib
sop += 0.5*omega_syms[k_mode] * (pp_ops[jsite] + qq_ops[jsite])
# H_G1
if k_mode in g1_data:
coeffs = g1_data[k_mode]
mat_g1 = np.diag(coeffs) * ev_to_au
sop += OpSite(f"V_G1_{k_mode}", 0, value=mat_g1) * q_ops[jsite]
# H_G3
elif k_mode == 5: # index 5 = 10a
mat_g3 = np.array([[0.0, g3_val],[g3_val, 0.0]]) * ev_to_au
sop += OpSite(f"V_G3_{k_mode}", 0, value=mat_g3) * q_ops[jsite]
# H_G2
for (k_mode, l_mode), coeff in g2_data.items():
j_1site = k_mode + 1
j_2site = l_mode + 1
mat_g2_current = np.diag(coeff) * ev_to_au
sop += OpSite(f"V_G2_{k_mode}_{l_mode}", 0, value=mat_g2_current) * q_ops[j_1site] * q_ops[j_2site]
# H_G4
for (k_mode, l_mode), coeff in g4_data.items():
j_1site = k_mode + 1
j_2site = l_mode + 1
mat_g4 = np.array([[0.0, 1.0], [1.0, 0.0]]) * (coeff * ev_to_au)
sop += OpSite(f"V_G4_{k_mode}_{l_mode}", 0, value=mat_g4) * q_ops[j_1site] * q_ops[j_2site]
sop = sop.simplify()
5.Setup MPO
[9]:
am_pot = AssignManager(sop)
_ = am_pot.assign()
pot_mpo = am_pot.numerical_mpo(subs=subs)
6.Setup Hamiltonian
[10]:
potential = [
[{tuple((k, k) for k in range(nsite)): TensorOperator(mpo=pot_mpo)}]
]
H = TensorHamiltonian(
ndof=len(basis), potential=potential, kinetic=None, backend=backend
)
operators = {"hamiltonian": H}
7.Setup initial states
[11]:
model = Model(basinfo=basinfo, operators=operators)
model.m_aux_max = 20
S2_state = [0.0, 1.0]
nprim_boson = basis[1].nprim if len(basis) > 1 else 10
boson_site = [1.0] + [0.0] * (nprim_boson - 1)
initial_weights = []
for isite in range(nsite):
if isite == 0:
initial_weights.append(S2_state)
else:
initial_weights.append(boson_site)
model.init_HartreeProduct = [initial_weights]
8.Execution
[12]:
jobname = "pyrazine"
simulator = Simulator(jobname=jobname, model=model, backend=backend, verbose=2)
simulator.propagate(
maxstep=1500,
stepsize=0.1,
reduced_density=(
[(0, 0)],
10,
),
energy=True,
autocorr=True,
observables=False,
observables_per_step=10
)
00:08:05 | INFO | Log file is ./pyrazine_prop/main.log
00:08:05 | INFO | Wave function is saved in wf_pyrazine.pkl
00:08:05 | INFO | The time unit in the netCDF file is fs
00:08:05 | INFO | Start initial step 0.000 [fs]
00:08:07 | INFO | End 0 step; propagated 0.100 [fs]; AVG Krylov iteration: 6.76
00:10:27 | INFO | End 100 step; propagated 10.100 [fs]; AVG Krylov iteration: 6.76
00:12:48 | INFO | End 200 step; propagated 20.100 [fs]; AVG Krylov iteration: 6.76
00:15:07 | INFO | End 300 step; propagated 30.100 [fs]; AVG Krylov iteration: 6.96
00:17:34 | INFO | End 400 step; propagated 40.100 [fs]; AVG Krylov iteration: 6.96
00:19:52 | INFO | End 500 step; propagated 50.100 [fs]; AVG Krylov iteration: 6.96
00:22:11 | INFO | End 600 step; propagated 60.100 [fs]; AVG Krylov iteration: 6.96
00:24:29 | INFO | End 700 step; propagated 70.100 [fs]; AVG Krylov iteration: 6.96
00:26:51 | INFO | End 800 step; propagated 80.100 [fs]; AVG Krylov iteration: 6.96
00:29:13 | INFO | End 900 step; propagated 90.100 [fs]; AVG Krylov iteration: 6.96
00:31:33 | INFO | Saved wavefunction 99.900 [fs]
00:31:36 | INFO | End 1000 step; propagated 100.100 [fs]; AVG Krylov iteration: 6.96
00:33:53 | INFO | End 1100 step; propagated 110.100 [fs]; AVG Krylov iteration: 6.96
00:36:17 | INFO | End 1200 step; propagated 120.100 [fs]; AVG Krylov iteration: 6.96
00:38:35 | INFO | End 1300 step; propagated 130.100 [fs]; AVG Krylov iteration: 6.96
00:40:56 | INFO | End 1400 step; propagated 140.100 [fs]; AVG Krylov iteration: 6.96
00:43:15 | INFO | End 1499 step; propagated 149.900 [fs]; AVG Krylov iteration: 6.96
00:43:15 | INFO | End simulation and save wavefunction
00:43:15 | INFO | Wave function is saved in wf_pyrazine.pkl
[12]:
(0.08830243567021843, <pytdscf.wavefunction.WFunc at 0x119813160>)
9.Plot (population, absorption spectrum)
[13]:
with nc.Dataset("pyrazine_prop/reduced_density.nc", "r") as file:
density_data_real = file.variables[f"rho_({0}, {0})_0"][:][
"real"
]
density_data_imag = file.variables[f"rho_({0}, {0})_0"][:][
"imag"
]
time_data = file.variables["time"][:]
[14]:
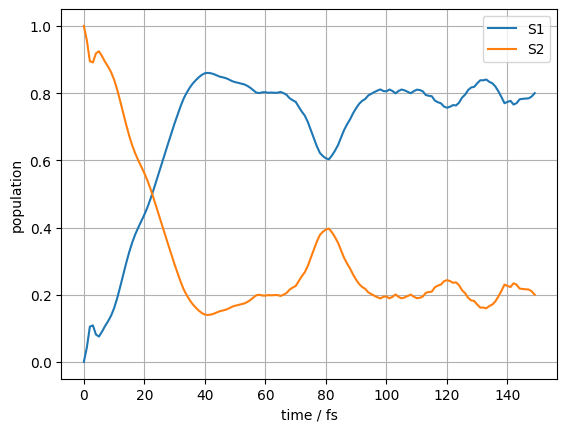
plt.ylabel("population")
plt.xlabel("time / fs")
plt.plot(time_data, density_data_real[:, 0, 0], label="S1")
plt.plot(time_data, density_data_real[:, 1, 1 ], label="S2")
plt.grid()
plt.legend()
plt.show()
[15]:
# omega : taken from https://pubs.aip.org/aip/jcp/article-abstract/110/2/936/475747
omega = [
0.0739, 0.1258, 0.1525, 0.1961, 0.3788, # Ag (6a, 1, 9a, 8a, 2)
0.1139, # B1g (10a)
0.0937, 0.1219, # B2g (4, 5)
0.0873, 0.1669, 0.1891, 0.3769, # B3g (6b, 3, 8b, 7b)
0.0423, 0.1190, # Au (16a, 17a)
0.1266, 0.1408, 0.1840, 0.3734, # B1u (12, 18a, 19a, 13)
0.1318, 0.1425, 0.1756, 0.3798, # B2u (18b, 14, 19b, 20b)
0.0521, 0.0973 # B3u (16b, 11)
]
E_0 = 0.5*sum(omega)-(3.94+4.89)/2 #experimental data (eV) : taken from https://doi.org/10.1063/1.466618
time, autocorr = pytdscf.spectra.load_autocorr(
"pyrazine_prop/autocorr.dat"
)
def h(t,tau = 150):
return np.exp(-np.abs(t)/tau)
freq, intensity = pytdscf.spectra.ifft_autocorr(time, autocorr * h(time), E_shift=E_0,window="cos")
# (cm^-1) -> (nm)
mask = freq > 0
wavelength_nm = 1e7 / freq[mask]
intensity_masked = intensity[mask]
plt.figure(figsize=(6, 4))
plt.plot(wavelength_nm, intensity_masked/ np.max(intensity_masked), color='red', lw=2)
plt.xlabel('wavelength / nm')
plt.ylabel('intensity')
plt.title('absorption spectrum')
plt.xlim(220, 280)
plt.grid(True, linestyle='--', alpha=0.6)
plt.show()
delta_t: 0.100[fs], max_freq: 333564.1[cm-1]